Your tools. Your data. One interface.
Nanome connects to the scientific software and databases your team already uses. MARA orchestrates them in plain English. Nanome XR puts the results in three dimensions. And Claude Code can drive the whole thing through our MCP server and skill.
Compatible with the platforms your team already relies on
Schrödinger LiveDesign
Pull Live Reports into XR. Iterate on molecular designs spatially.
CDD Vault
Query saved searches and find similar molecules by SMILES.
OpenEye / Cadence
FastROCS shape search, MD trajectory parsing, cryptic pocket evaluation
Discngine 3decision
Browse and retrieve structures from 3decision projects.
KNIME Server
Trigger KNIME workflows, check job status, analyze results

CCG MOE
Import MOE session files directly into Nanome.
Boltz
Biomolecular structure prediction and analysis

OpenFold
Open-source protein structure prediction workflows
Public databases and open-source tools
0+ scientific tools. One conversation.
MARA ships with a growing library of built-in tools across 26 categories. Docking, structure prediction, cheminformatics, genomics, materials science, and more. Run them in plain English or chain them into multi-step workflows.
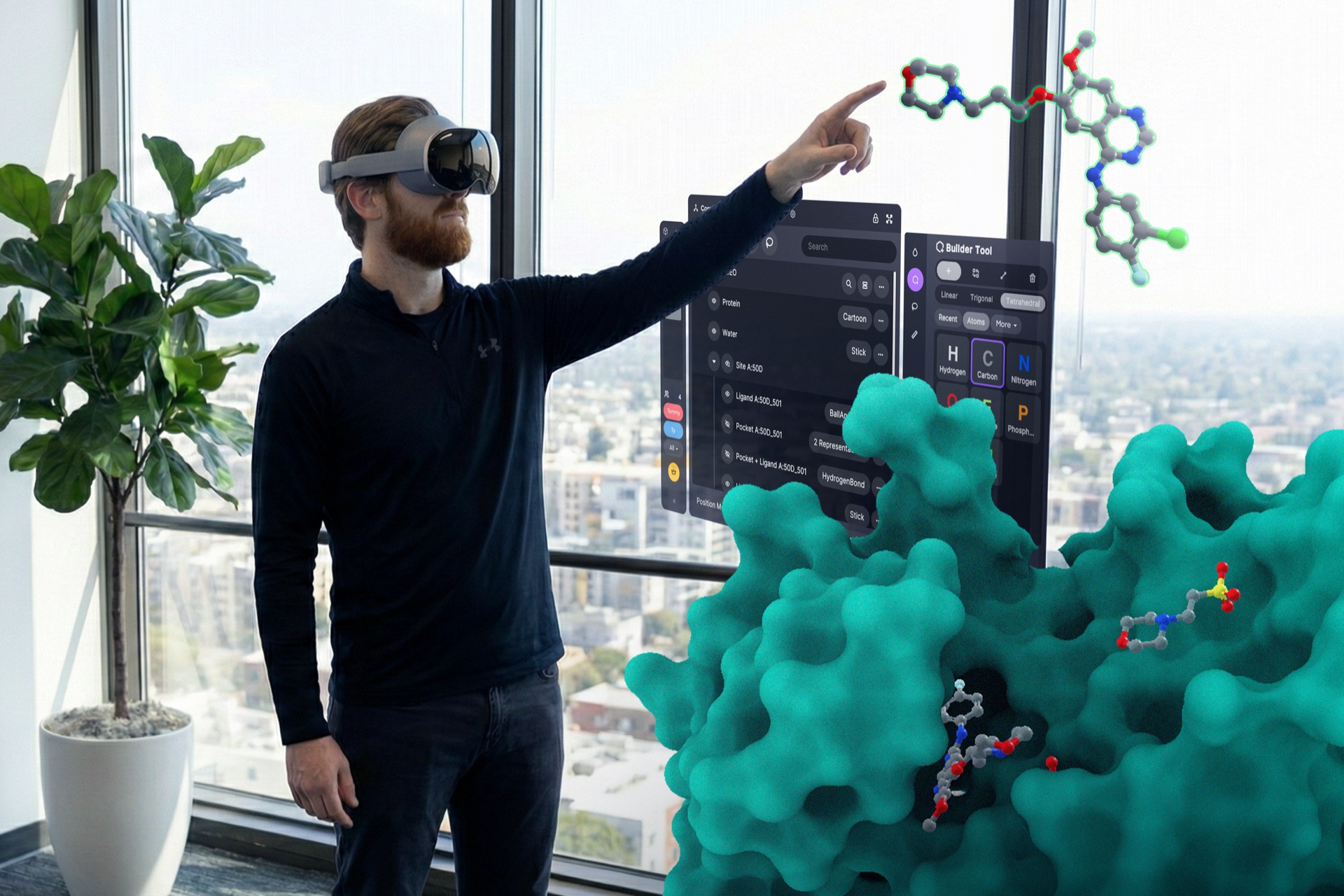

Build custom tools. Wire in your own infrastructure.
Python Snippets
Drop in a snippet, MARA spins up the sandbox. Use AI to write the snippet if you want.
HTTP Endpoints
Point MARA at any REST API and it becomes a conversational tool.
SQL Queries
Query your databases through MARA. Results flow into the rest of your analysis.
Enterprise teams have built integrations with proprietary scoring functions, internal databases, and custom analysis pipelines. Whatever you build is instantly available across your org. Learn how to create tools →
Programmatic access. AI-native workflows.
Jupyter notebook integration
Call MARA from Jupyter. Install the Python package, drop in your API key, and run any MARA tool from the same notebook you're already using.
Learn more →Claude Code skill and MCP server
Nanome publishes a Claude Code skill and an MCP server. Coding agents can prep config files, run docking, parse results, and reach the full MARA tool library directly.
Learn more →REST API
Every MARA tool is a REST endpoint. Drop Nanome's capabilities into your own pipelines, CI/CD, and automation infrastructure.
Explore documentation →All of this, from inside XR.
Every integration, every tool, every workflow is reachable from inside Nanome XR. Run docking, query databases, fire off workflows, walk through the results spatially, all without taking the headset off. MARA voice commands let you do it hands-free.
 RCSB PDB
RCSB PDB